Mendelian
inheritance
Alleles:
What are they?
Sex
determination
X-linked
traits
Pedigree
analysis
Assumptions
of Mendelian Inheritance
-
Constant
environment (temperature,
nutrition, sunlight etc.)
-
Traits
only influenced
by known gene loci
-
Genes
assort independently
-- zero "linkage"
Mendelian
inheritance
Use
Flowers
tutorial: p:\data\biology\biol14\tutorial\flowers.exe
Know
these terms:
-
Gene
Locus
-
Gene
Product
-
Allele
-
Dominant
and Recessive
alleles; Null alleles
-
Trait
-
Genotype
-
Phenotype
-
Hybrid:
Monohybrid;
Dihybrid -- Dihybrid Cross
-
Self
cross
-
Test
cross or Back
cross (when are these the same -- and when not?)
-
Probability
rules
Problems:
Mendelian
Genetics Practice from
a course at MIT.
Other
links:
Mendel's Peacocks
Horse
Genetics: describes interesting horse phenotypes and genotypes,
crosses and results
Online Mendelian Inheritance in Man: a
professional physician’s reference on inherited diseases
Alleles:
What are they?
An allele
is
a particular version of a given DNA sequence. "Allele" is a
relative
term, implying more than one possible version or copy, like different
editions
of a book. Like editions of a book, all existing alleles result
from
a process of change, either gradual or drastic change.
Examples:
Allele 1
ATCGTTAGATTACAGATTTACCGA
TAGCAATCTAATGTCTAAATCCGT
Allele 2
ATCGTTAGATTCCAGATTTACCGA
TAGCAATCTAAGGTCTAAATCCGT
Allele 3
ATCGTTAGTGTAATGATTTACCGA
TAGCAATCACATTACTAAATCCGT
Allele 4
ATCGTTAG-GATTTACCGA
TAGCAATC-CTAAATCCGT
Notice that
there can
be more than two
possible alleles for a given gene locus (but only
two at a time, in a given diploid individual.)
Multiple
alleles
can mean many different possible combinations for
individuals.
An example of multiple alleles is human blood type -- A or B alleles
encode
a blood serum protein, whereas the O allele makes no protein ( a null
allele.)
Gene loci which confer traits of tissue type may have 20 or more
different
alleles.
Natural and
"artificial"
alleles
-
Natural
alleles result
from evolution, the process of natural selection.
-
Artificial
alleles can
be created by molecular genetics.
-
Both
natural and artificial
alleles can be used by the scientist for breeding purposes.
Alleles can be
observed
as DNA polymorphisms, using restriction digest and gel electrophoresis
(see Week 7).
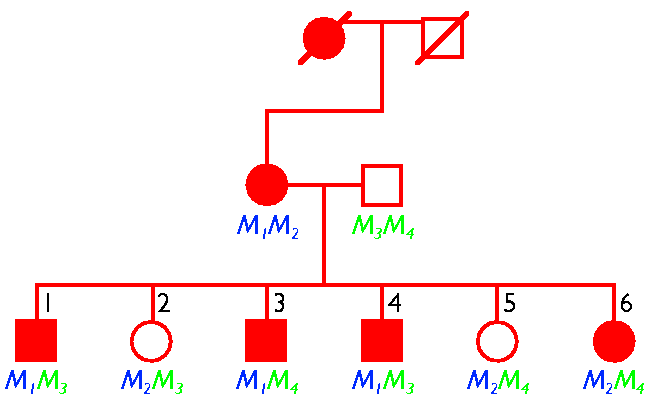
An allele may
be linked
to an inherited disease--a clue as to the gene locus defective in the
disease.
Which of
the
four alleles (M1-M4) is linked to this disease?
Is the disease
likely
to be dominant or recessive?
For a
Kenyon student's
report on an inherited disease in her family, see Tuberous
Sclerosis
Complex.
Alleles confer
traits,
by
expressing gene products, which are either mRNA and protein, or
a functional RNA. But how they determine a "visible
trait" is not simple. Consider this:
Problem:
Fruit
fly eyes have two pigments, brown and scarlet. Normal flies make
both
pigments, but a strain with defective gene B has brown eyes, and a
strain
with defective gene S has scarlet eyes.
In
the wild type, WHICH GENE (B or S) makes which pigment (brown or
scarlet)?Solution
In
practice, the most
common "new" alleles (arising out of mutation) are often named for a
phenotype
resulting from the absence of their gene product. Thus the allele
for a gene producing scarlet pigment is named "brown" for the brown eye
in the absence of scarlet pigment.
Consider
albinism,
or loss of pigmentation, a very common phenotype observed in many
species
of animals and plants. Alleles can confer loss of pigmentation in
two different ways:
-
Recessive
albinism.
The allele encodes an enzyme which converts pigment precursors into
dark
pigment; or a protein required for pigment deposition. (Humans;
mice;
penguins.)
-
Dominant
color suppression.
The allele encodes a regulatory protein which represses synthesis or
deposition
of pigment. (Horse; foxglove)
Traits are not
actually
inherited like "beads on a string." Traits result from complex
interactions
(1) among the products of genes; (2) between genes and regulatory
proteins
expressed by other genes; (3) between genes and proteins, and
environmental
factors such as nutrients, temperature, etc.; (4) chance effects during
development.
The reason
Mendelian
inheritance can be seen to "work" is that in many cases we can hold all
the above factors constant, for a given genotype (genes
affecting
a trait) and a given
phenotype (appearance of trait).
Other ways
alleles
can work (still at a SINGLE gene locus)
-
Codominance
or incomplete
dominance. Codominant alleles each contribute to the
phenotype;
for example A and B blood type alleles together produce AB blood
type.
Incomplete dominance means that the hybrid produces a lesser degree of
the dominant phenotype than the purebreeding dominant.
-
Lethality.
If an allele (either dominant or recessive) results in death before
birth,
a class of progeny will be absent from the offspring. What ratios
will result?
Pleiotropy.
One
allele (or pair of recessive alleles) at one gene locus can result in
many
diverse effects throughout the body.
Artificial
alleles.
Today
we can use molecular genetics to create artificial alleles in
transgenic
animals and plants. There are two ways to make a transgene,
which have different consequences for inheritance:
-
Allele
insertion.
Inject or transfect DNA into a fertilized egg. The DNA sequence
gets
taken up somewhere in the genome, but not at the same position
as
any homologous gene. The position on the gene map is
unpredictable
(but fixed once the allele is inserted.)
-
Allele
replacement.
A DNA sequence containing a linked selective marker (such as drug
resistance)
is put into embryonic stem cell culture (ES cells). The ES cells
are put into a blastula, which develops into a chimeric offspring;
eventually
a pure-breeding line is bred. In this case, the new allele recombines
homologously and replaces an allele at a standard map position.
Dihybrid Cross
Advanced
Problem. Explain how to breed transgenic mice to create a mouse
model
for sickle-cell anemia.
Sickle
Cell Anemia.
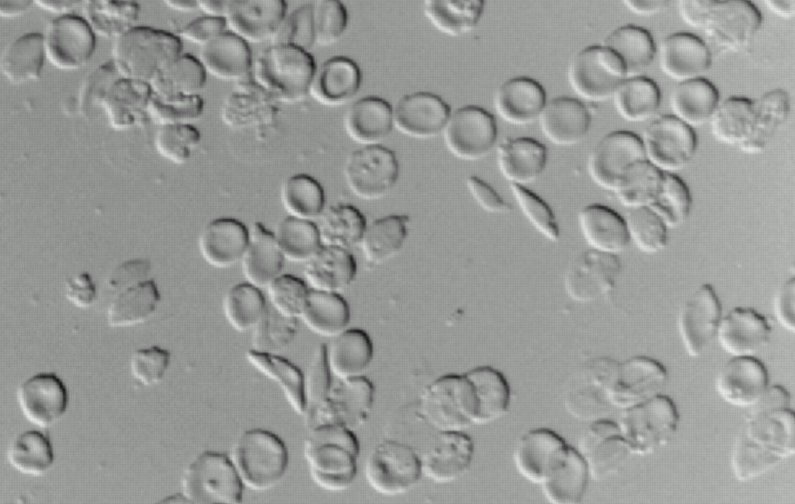
Pászty et al,
Science
1997 October 31; 278: 876-878.
Sickle-cell
disease
is caused by a single base pair defect in human beta-globin.
-
Double-recessive
genotype--red
blood cells sickel under stress.
-
Single-recessive
heterozygote--cells
sickle only when attacked by malaria parasites. Prevents malaria.
To test drug
therapies
for sickle disease, can we generate a transgenic mouse model for human
sickle-cell anemia?
For the
model strain
to exhibit sickle-cell pathology, the native mouse genes--all at
separate
loci--must be defective (null alleles.) We have a transgenic
mouse
strain, containing
human Hb-alpha, Hb-beta-sickle
on a transgene,Tg(Hu), inserted
somewhere
in the mouse genome (not at the mouse globin genes.) But the
mouse
still has its own genes producingalpha
and
beta globin.
To
construct this strain,
the transgenic strain was intercrossed with a mouse strain heterozygous
for null alleles for alpha
and
beta globin.
Tg(Hu)
Mouse-alpha-Hb
Mouse-beta-Hb
--------
----------------------
----------------------
Tg(Hu)
Mouse-alpha-Hb
Mouse-beta-Hb
X
-- Mouse-alpha-Hb
Mouse-beta-Hb
--------
--------------------
--------------------
-- ----
----
How many
generations
would you need to cross?
What
proportion of
mice would show the desired phenotype of blood with entirely human
globins?
Tg(Hu)
----
----
------------
------------------ -------------------
Tg(Hu)
----
----
What would
the researchers
have to do to create a similar model with exclusively normal human
blood?
Why would this be important in order to use the model? (For
further
interest, read Ryan et al, 1997.)
Sex
determination
What is the
function
of sex? Why have so many animal and plant species evolved the
complex
mechanisms of sexual recombination?
Natural
selection favors
diverse combinations of traits, because when the environment
changes,
there is greater chance that some individuals will survive.
How do
animals and
plants develop two different sexual types (sexes)? Different
species do it differently:
-
Male and female organs
develop
on the same individual. (Garden pea plant; invertebrate worms)
-
Juvenile is born
female, later develops
into male; or vice versa. (Some fish and other vertebrates)
-
Diploid (female) versus
haploid,
from unfertilized eggs (male) (Ants, bees, other social
insect
colonies)
-
Sex chromosomes--X, Y
(mammals) or
W, Z (birds; moths). One member of pair (male Y, or female W)
is largely degenerate, having lost most of its genes through
evolution.
(Why?)
Read
the latest about evolution of X and Y.
X-linked
inheritance
Species
that show X,
Y sex determination can have two different mechanisms of addressing gene
dosage:
-
Random
inactivation of one X or the other, in early embryonic cells.
(Humans)
-
Half
down-regulation of gene expression from both X chromosomes. (Drosophila)
In
either case, traits encoded by genes on the X chromosome will show
X-linked
inheritance. A female carrying two recessive X-linked alleles,
when
crossed with a wild-type male will produce criss-cross
inheritance. This is because the
Y chromosomes from the male all behave as null alleles
(Why?).
So the recessive allele from the female parent is always expressed in
the
male offspring. But her female offspring will receive one
wild-type
allele from the father.
The other,
normally
paired chromosomes are called autosomes.
For
X-linkage, you need to know the results of these crosses:
A
A
a
X X with X
Y
----> offspring?
a
a
A
X X with X
Y
----> offspring?
A
a
a
X X with X
Y
----> offspring?
A
a
A
X X with X
Y
----> offspring?
You
also need to FIGURE OUT THE PARENTS of a given combination of offspring.
X-linked
traits
X-linked
traits are
particularly common because they only need one recessive allele present
for the phenotype to be expressed in the male. Some examples:
-
Colorblindness.
About
a
third
of all men
are partly color-blind.
Defective alleles are so common because (a) their effect is non-lethal,
and (b) the genes for red and green photoreceptors are extremely
similar
and lie close together on the X chromosome, where they can recombine
(cross
over) with each other by mistake. Sometimes a color-blind person
can see a different "hybrid" color that no one else can!
-
Duchenne
Muscular Dystrophy
One of the
interesting
features of Fragile X syndrome is the role of imprinting
by methylated genes. (See next week.)
Pedigree
analysis
Problems: More
Pedigrees, here ; SOLUTIONS;
Still more pedigrees **Medical Pedigrees
To
understand "dominant" and "recessive":
-
A
Dominant gene makes MUCH MORE than enough protein to cause a trait.So
only
one is needed; perhaps only in some cells.
-
An
Incompletely Dominant gene makes BARELY ENOUGH protein for the
trait.
So TWO COPIES are needed for the FULL trait.
-
A
Recessive gene makes no protein; inactive or partly active protein; or
not enough protein for the trait.
Partial
Penetrance and Variable Expressivity
In real
pedigrees of
real people, inheritance of any trait (dominant or recessive) is often
confounded by partial penetrance or by variable
expressivity
of a trait.
-
Penetrance
is the
percentage of individuals with a genotype who actually show the
trait.
If only 80% of people with the genotype actually develop the trait,
then
you could pass on a trait without showing it -- even if the trait is
"dominant"!
-
Expressivity
is the degree of the trait. For example a genetic defect causing
mental retardation (such as Fragile X) can result in individuals with a
very wide range of intellect; and you cannot predict the degree of
expression.
|


