Biology Dept Kenyon College |
|
 |
Biology Dept Kenyon College |
|
 |
|
1. A fly of wild-type phenotype (red eyes, full
wings) is crossed to a wingless fly with brown eyes. The progeny
are:
Estimate the map distance between the eye color and wingless
traits. Show your work.
Crossover frequency = (7 + 8)/(62+61+7+8)
= 0.108 = 10.8% 2. Two black mice are crossed, producing several dozen progeny. The color ratio of the progeny is approximately 9 black: 4 white: 3 brown. Explain these results, defining alleles for the genes, and show an enzyme diagram. It works the same as the beetles: 
If A is missing, no brown can be made; mice are white, regardless of B (so 1 + 3 = 4, four white). If B is missing, but A is present, mice are brown (3 brown). 3. An eyeless female fly is crossed to a male fly with normal eyes. Among the progeny, the males all lack eyes, and the females all have shrunken heart-shaped eyes. Explain. Eyeless trait is X-linked. The male progeny have to get their one X from the eyeless mother, so they are all eyeless. The females get one normal X from the father and one eyeless X from the mother. They show an intermediate phenotype (shrunken heart-shaped) because the trait is codominant. Because males have the XY, they can only show the phenotype from one X. In humans, males never show codominance for X-linked traits. 4. In Flowers, a blue flower is crossed with a pink flower; the progeny are half turquoise (light blue), and half pink. Explain. The blue hides a recessive allele (BO). The pink is pure-breeding (PP). The progeny are PB (turquoise/light blue) or PO (pink). 5. Chemical analysis of the genome of a newly discovered microbe shows 10% adenine. State the percentage of each of the other DNA bases. 10% A = 10% T 6. Compare and contrast the organization of gene information in bacteria and in eukaryotes. Bacterial genomes are highly compact, with 95% coding content. Many genes are organized in operons (tandem series of genes with related function, transcribed off one promoter). Each gene is intact, encoding one RNA or protein sequence. There are occasional interuptions by insertetd viral genomes and transposable elements. Eukaryotic genomes resemble those of bacteria, in that they are composed of DNA, and contain genes as discrete regions of sequence encoding linear information for RNA or protein products. However, eukaryotic gene sequences are interspersed with intergenic non-coding sequences, and are interrupted with non-coding sequences called introns. About 5% of the genome appears to encode product (RNA or protein). Overall, 45% of the human genome has originated from viral or transposable elements. 7. Draw the structure of deoxy-guanine nucleotide monophosphate (without looking). How would the RNA nucleotide differ? Here are all of the nucleotides:
8. Identify this molecule. What are its parts, and how does it function in cells? 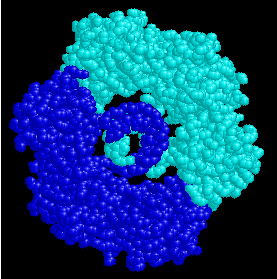 DNA Polymerase III Beta clamp -- two beta subunits clamped around a DNA helix. The DNA helix is being replicated; Pol III synthesizes the new DNA strand.
9. State the steps of DNA replication. Propose what might happen to the cell at each step, if the step fails, or if the needed enzyme is absent.
10. What is the Environmental Genome Project? How does it differ from the main Human Genome Project? 11. Is diabetes an inherited disease, or an environmental condition? Explain. Both. Certain alleles of certain
genes cause predisposition to diabetes. However, environmental
factors such as diet determine the age of onset and severity of the
disease. Different imprinting (methylation of DNA, or methylation, acetylation, phosphorylation of histones) in the mother and father. 13. What is the histone code? Explain how histones interact with genetic information. The histone code involves chemical modifications to DNA-associated histones (or to DNA itself, as in the case of methylation.) Acetylation of histone usually turns a gene on (enables transcription to make messenger RNA). Methylation of histone (or of DNA itself) usually turns a gene off. Phosphorylation may either enhance or repress gene expression. Thus, the penetrance or expressivity of an allele could be affected by histone modification. 14. Explain how the Y chromosome evolved in humans. In birds, how and why did the sex-determining chromosome evolve differently? The original sex-determination gene was located on an autosome. Since every mating event required a parent with one (but only one) copy of this sex-determining autosome, there was no selection against deleterious genes on this chromosome--especially those closely linked to the sex-determining gene. So over the generations, the sex-linked genes were lost by reductive evolution. As the chromosome shrunk, more genes became sex-linked, and they too became lost on what is now the Y chromosome. In birds, the sex-determining gene happened to be on a different autosome, and it happened to determine female sex, not male. So a different chromosome underwent reductive evolution. |