S. cerevisiae Prp8 Active Site of Spliceosome
Beija Waltmunson '27, Ava Merickel '27
View Type:
Contents:
I. Introduction
II. General Structure
III. Prp8 Interaction With Aar2
IV. Prp8 Interaction With the Spliceosome
V. Real-world Applications
VI. References
I. Introduction
Prp8 is a large protein component of the U5 small nuclear ribonucleoprotein (snRNP), a structural and catalytic component of the spliceosome. The spliceosome is a multi-megadalton ribonucleoprotein (RNP) complex that converts precursor mature RNA sequences into mature RNA through the removal of non-coding intervening sequences, or introns. This splicing process produces viable sequences of RNA for translation and is essential for RNA processing.
Prp8 plays a critical role in the formation of the U5 catalytic core and its ability to perform RNA processing. Prp8 facilitates the positioning of splice site consensus sequences at intron boundaries with high fidelity via crosslinking to the U5, U6, and substrate residues on pre-mRNA. In the cytoplasm, Prp8 also interacts with Aar2. Aar2 is a cytoplasmic U5 assembly factor that is necessary for spliceosome formation and is later replaced by Brr2 to activate the complex for pre-mRNA splicing. This crystal structure reveals Prp8 in complex with the full-length Aar2 protein.
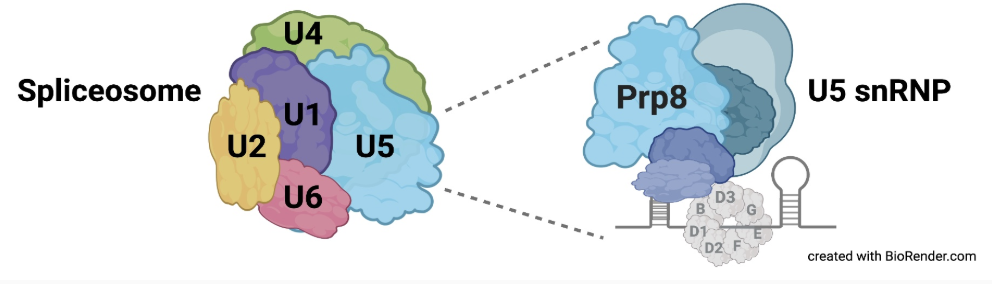
Fig. 1. Prp8 in complex with U5 (Will and Lührmann 2011)
II. General Structure
Prp8 has four domains: the Jab1/MPN domain, the RNaseH-like domain, the reverse transcriptase-like domain, and the endonuclease domain.
The large domain
contains the reverse transcriptase-like domain
and a
small type II restriction endonuclease-like domain
which are connected by a
linker
. The reverse transcriptase-like domain is composed of two significant subdomains: the palm/fingers
and the thumb
which form a deep cleft that resembles the nucleic acid template and primer in polymerases.
The palm subdomain is made up of 4 conserved motifs, each containing a catalytically important residue, motif A, B, and C. The palm sub-domain forms a four-stranded antiparallel beta sheet
flanked by
alpha helices , which house the catalytic amino acid residues essential for kinase activity and phosphorylation .
III. Prp8 Interaction With Aar2
Aar2 is a A1 mediates the formation of pre-U5 snRNPs in the cytoplasm. The domain architecture of Aar2 is comprised of the N-terminal-domain and the
C-terminal-domain . To mediate U5 assembly, Aar2 facilitates the folding back of the RNase-like and Jab1/MPN domains so they can interact with the large domain. The C-terminal tail of Aar2 is crucial to bring the RNaseH-like and Jab1/MPN domains together. The C-terminal tail of Aar2 extends through the cleft between the reverse transcriptase fingers and thumb subdomains of Prp8. It interacts with Prp8 through hydrophobic interactions and . In addition, Aar2s C-terminal end forms a parallel β-sheet, zipping together the β-hairpin of the RNaseH-like domain and the β-barrel of the Jab1/MPN domain . By binding with the Jab1/MPN domain, Aar2 prevents Brr2, a helicase that ultimately activates pre-mRNA splicing, from binding. Aar2 occupies this domain until the pre-snRNP enters the nucleus. Phosphorylation of triggers refolding and Aar2 is released and Brr2 binds to complete U5 snRNP biogenesis.The β-hairpin is also thought to be involved in protein-protein interaction responsible for positioning the RNaseH-like domain in the U5 snRNP part of the splicosome.
IV. Prp8 Interaction With the Spliceosome
Prp8 Interaction with U5:
Interactions between U5 and Prp8 help position the pre-mRNA in the 5’ splice site for high fidelity splicing. Crosslinks lie within the small type II restriction endonuclease-like domain of Prp8 and interact with the exon binding loop 1 of the U5 snRNA (U5 SLI). This loop interacts with the 5’ exon of pre-mRNA to initiate splicing. Four highly conserved lysines,
are located in the region that surrounds the U5 SLI as it interacts with the 5′ exon of pre-mRNA. Lysine residues maintain a positive charge and often play important roles in stabilization of specific conformations of the phosphate backbone of RNA.
Prp8 Interaction with pre-mRNA: Additionally, Prp8 is involved in a functional interaction with both splice sites. Before splicing begins, the 5’ splice site (5′SS) GU dinucleotide of pre-mRNA forms a highly specific crosslink to Prp8 residues
to properly position the spliceosome for activity. During the second catalytic step of splicing, Prp8 forms crosslinks with the 3’ splice site (3’SS) and the branch point of pre-mRNA to facilitate the release of the intron and ligation of the two exons. Crosslinks to the 3’SS and the branch point of pre-mRNA occur via Prp8 residues
1503 through 1673 located within the linker domain and the restriction endonuclease-like domain.
V. Real-world Applications
The structure of Prp8 provides insight into how the spliceosome evolved. The conserved structural and sequence similarities between the reverse transcriptase-like domain and the Th/X domain share resemblance with their counterparts in bacterial and fungal group II intron encoded proteins. This high conservation of the Prp8 protein is due to the important role that it plays in maintaining accurate gene transcription and gene regulation through splicing. Thus, the disruption or absence of Prp8 leads to the development of physiological and pathological diseases including cancers. Mutations to the Prp8 can induce abnormal splicing events such as intron retention and exon skipping that give rise to isoforms that can cause tumorigenesis, metastasis, and drug resistance. Due to its role in binding both mRNA and U5 and U6, Prp8 can impact spliceosome assembly and therefore reduces the number of functional spliceosomes, consequentially increasing the occurrence of splicing abnormalities. In cancer cells, mutations can alter Prp8 function. Most of these mutations are located in the C-terminal Jab1/MPN domain . Prp8 dysfunction not only affects metastasis and tumor growth but also enhances tumor invasiveness by regulating signaling pathways such as FAK/AKT, PI3K/Akt, and p53 pathways and their critical factors. Furthermore, Prp8 is capable of inducing apoptosis through the p53 pathway. This presented potential for Prp8 use in cancer treatment. This research is currently in the experimental stage for tumors that exhibit resistance to conventional treatments.
VI. References
Che-Sheng Chung, Chi-Kang Tseng, Yung-Hua Lai, Hui-Fang Wang, Andrew J Newman, Soo-Chen Cheng, Dynamic protein–RNA interactions in mediating splicing catalysis, Nucleic Acids Research, Volume 47, Issue 2, 25 January 2019, Pages 899–910, https://doi.org/10.1093/nar/gky1089
Fernández-Espartero CH, Rizzo A, Fulford AD, Falo-Sanjuan J, Goutte-Gattat D, Ribeiro PS. Prp8 regulates oncogene-induced hyperplastic growth in Drosophila. Development. 2018 Nov 12;145(22):dev162156. doi: 10.1242/dev.162156. PMID: 30333215; PMCID: PMC6262796.
Grainger, R. J., & Beggs, J. D. (2005). Prp8 protein: at the heart of the spliceosome. RNA (New York, N.Y.), 11(5), 533–557. https://doi.org/10.1261/rna.2220705
Huang G, Wang D, Xue J. Research Progress on the Relationship Between PRPF8 and Cancer. Curr Issues Mol Biol. 2025 Feb 26;47(3):150. doi: 10.3390/cimb47030150. PMID: 40136404; PMCID: PMC11941625.
MacRae AJ, Mayerle M, Hrabeta-Robinson E, Chalkley RJ, Guthrie C, Burlingame AL, Jurica MS. Prp8 positioning of U5 snRNA is linked to 5' splice site recognition. RNA. 2018 Jun;24(6):769-777. doi: 10.1261/rna.065458.117. Epub 2018 Feb 27. PMID: 29487104; PMCID: PMC5959246.
Preussner, M., Santos, K. F., Alles, J., Heroven, C., Heyd, F., Wahl, M. C. & Weber, G. (2022). Structural and functional investigation of the human snRNP assembly factor AAR2 in complex with the RNase H-like domain of PRPF8. Acta Cryst. D78, 1373â1383.
Sun, Y., Zhang, L., Fang, Z., Liu, D., Shao, M., Liu, Y., Liao, B., & Jin, Y. (2023). PRPF8 controls alternative splicing of PIRH2 to modulate the p53 pathway and survival of human ESCs. Journal of Cellular Physiology, 238, 1909–1920. https://doi.org/10.1002/jcp.31066
Weber, G., Cristão, V. F., de L Alves, F., Santos, K. F., Holton, N., Rappsilber, J., Beggs, J. D., & Wahl, M. C. (2011). Mechanism for Aar2p function as a U5 snRNP assembly factor. Genes & development, 25(15), 1601–1612. https://doi.org/10.1101/gad.635911
Will, C. L., & Lührmann, R. (2011). Spliceosome structure and function. Cold Spring Harbor perspectives in biology, 3(7), a003707. https://doi.org/10.1101/cshperspect.a003707
Back to Top