Full-Length Human IFIT5
Ifti Rahman & Isaac Yu
Contents:
View Type:
I. Introduction
IFIT5 belongs to a class of proteins called the Interferon Induced proteins with Tetratricopeptide repeats (IFITs) which confer immunity against viral pathogens in response to cytokines (signaling molecules) known as Interferons. Four members of the IFIT protein family have been identified in humans thus far, the genes encoding which are found in a cluster on chromosome 101.
Expression of IFIT genes is negligible in healthy cells but is rapidly induced at large levels in the wake of a viral infection. While IFIT transcription is particularly stimulated by Type I Interferons, it can also be driven, albeit weakly, by Type II Interferons1.
IFITs act to inhibit or prevent synthesis of viral proteins by host ribosomes. While both IFIT1 and IFIT2 can achieve this through interacting with eukaryotic Initiation Factor 3 (eIF3), IFIT5 exerts its antiviral action by directly engaging viral RNA2. In particular, IFIT5 targets single stranded (ss) RNA bearing a 5’-triphosphate group (PPP-RNA), a potent molecular signature which identifies viruses as non-self or foreign within the body. IFIT5 can bind directly to PPP-ssRNA or double stranded (ds) PPP-RNA with a 5’ overhang of at least 3 nucleotides (nt) in a sequence-independent manner to inhibit viral protein translation.
II. General Structure
IFIT5 is a single polypeptide in length. It consists of 24 alpha helices, of which 18 form tetratricopeptide repeats (TPRs), characteristic of the IFIT family. Each TPR is a 34 amino acid helix-turn-helix motif which typically occurs in tandem repeats of 2-20 to yield a solenoidal architecture. The remaining 6 ɑ-helices of IFIT5 disrupt the regular repeating relationship between successive TPRs, producing made up of distinct TPR bundles. is made up of 2 TPRs capped off at either end by helices ɑ1 and ɑ2. These helices are linked together by a containing a Cys-His-Phe-Thr-Trp that is highly conserved among the IFIT family2. and contain 4 and 3 TPRs respectively and form a supercoil with each other. Subdomains II and III are connected to each other by a pair of extended helices, ɑ15 and ɑ16, which make a so that Subdomains II and III can supercoil against each other. This makes IFIT5 generally clamp shaped with a 28 Å by 15 Å pocket rich in positively charged residues that readily accommodates negatively charged nucleic acids2. The IFIT5 molecule depicted here is bound to a oligo-uracil
III. PPP-ssRNA Binding
IFIT5 binds to the 5’ end of PPP-ssRNA in a non-sequence-specific fashion. The 5’-triphosphate group, deep within the pocket, makes several with helix α2 (Gln 41) and the inner surface of Subdomain II (Lys 150, Arg 186, Tyr 250, and Arg 253) via hydrogen bonding and salt bridge interactions. An unknown metal ion appears to play a crucial role in the recognition of 5’-PPP through of Glu 33 and the α and γ phosphates of PPP-RNA2. Na+ is depicted here, but this ion is most likely Mg2+ as it is more commonly found as a metallic cofactor.
IFIT5 has specific interactions with the 5’ phosphates and 2’ hydroxyls of the first 3 nt. In particular, the 5’ phosphate of forms a hydrogen bond with Tyr 254 and the 5’ phosphate of forms a salt-bridge with Lys 257 and Arg 260 and a hydrogen bond with Gln 288. While the 2’ hydroxyls of and also make hydrogen bonds with Tyr 156 and His 287 respectively, these are not critical to the protein's RNA-binding ability2.
On the other hand, IFIT5's interactions with RNA nucleobases are exclusively non-specific. N1 makes with Gly 152, Gly 153, and Tyr 156, and stacks against N2, which further stacks against Phe 337 in Subdomain III. The remaining bases also stack against Phe 339 in a similar fashion as N1 and N22. Consequently, IFIT5 can accommodate any 5’-PPP-RNA, regardless of its sequence.
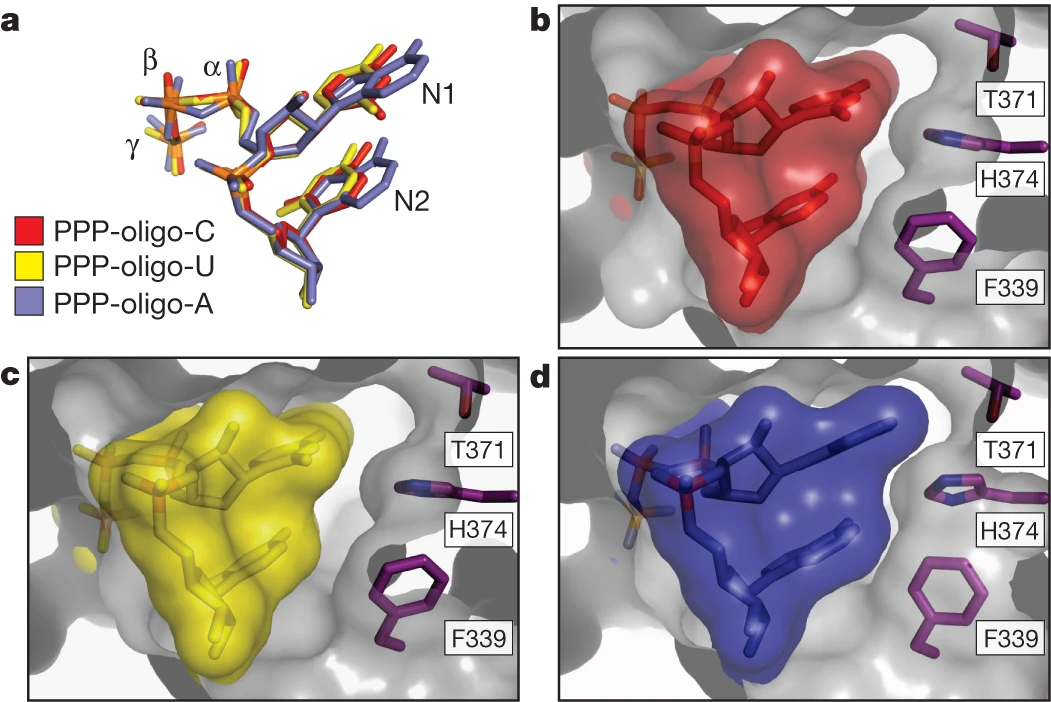
Figure 1. IFIT5 complexes with oligo-cytosine (b), oligo-uracil (b), and oligo-adenine (d) prepared by Abbas et al. highlighting sequence-independent binding of IFIT5 with 5'-PPP-RNA ligand. Larger purine bases like adenine (d) can make additional van der Waals contacts with IFIT5 at residues shown to stabilize interaction2.
The RNA binding pocket of IFIT5 is approximately 15 Å wide and, as such, can not fit dsRNA or capped RNA. The narrow groove readily binds ssRNA and can occasionally bind dsRNA, provided that it also contains a 5' overhang upwards of 3 nt. In its unbound state, IFIT5 appears to have a more open conformation to allow smooth entry of any ligand RNA. Upon binding, however, the protein adopts a more closed conformation whereby Subdomain III is forced closer to the rest of the molecule by considerable movement at the Pivot region2. This allows for enhanced protein-RNA interactions to occur.
IV. Roles in Immune Pathways
The 5'-triphosphate moiety is a common feature of the genomes of many negative sense ssRNA viruses such as Influenza A Virus (IAV) and Vesicular Stomatitis Virus (VSV). Positive sense ssRNA viruses with 5' caps also generate PPP-ssRNAs as transient intermediates while replicating3. Since RNAs in human cells do not typically bear a 5'-PPP, it is swiftly recognized as non-self and targeted by the innate immune system. IFIT5 represents a key player in that process, partaking in the first line of defense against viral pathogens.
Outside of directly engaging viral RNA, IFIT5 acts as a positive regulator in the classical NF-κB (Nuclear Factor-κB) signaling pathway4. The NF-κB family of inducible transcription factors are involved in a wide array of cellular processes ranging from inflammation to development of cancers, with new roles being discovered routinely. In absence of stimuli (cytokines, for instance), NF-κB remains inactivated through association with proteins of the inhibitory IκB family. As shown in Figure 2, IFIT5 facilitates recruitment of IκB kinase (IKK) to TAK1, resulting in the phosphorylation and subsequent degradation of IκB proteins. This leaves NF-κB (complex of proteins p50 and p65 shown in Figure 2) free to enter the nucleus and drive expression of target genes, which may be IFIT genes themselves4.
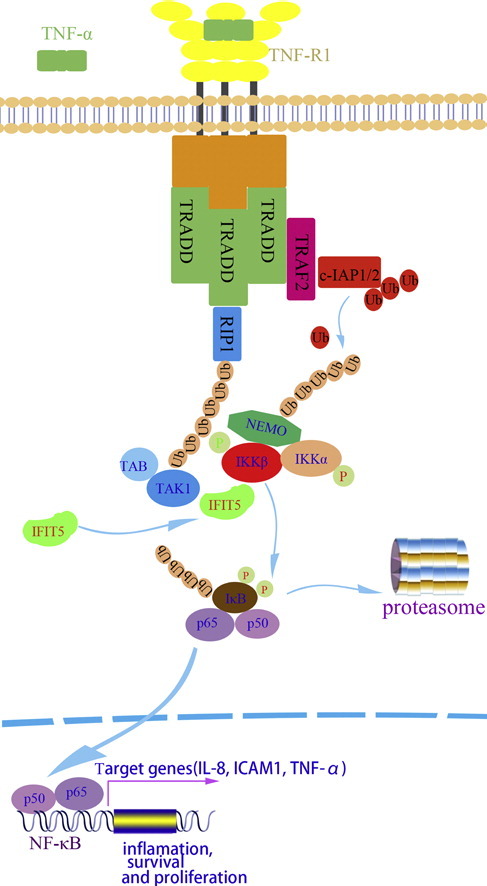
Figure 2. IFIT5 acts as a positive regulator of the canonical NF-κB signaling pathway4.
In recent years, there has been a growing interest in the role of IFITs, including IFIT5, beyond just antiviral activity, especially in cancer. It has been found that the IFIT5 gene can behave as an oncogene and IFIT5 is overexpressed in both prostate and bladder cancer cells5, 6. The ability of IFIT5 to bind any uncapped RNA sequence with a 5' overhang allows it to bind to endogenous precursor-microRNAs (pre-miRs) -363 and -99a in prostate and bladder cancer cells respectively. These miRNAs suppress the epithelial-mesenchymal transition (EMT) which prevents metastasis of cancer cells. However, after binding, IFIT5 recruits an exoribonuclease called XRN1 which degrades the pre-miRNAs, resulting in elevated EMT5, 6. The capability of IFIT5 to function as both an important antiviral and a cancer-promoting molecule sheds further light on the nuanced workings of the immune system and represents a necessary area of research.
V. References
1. Fensterl, V., & Sen, G. C. (2011). The ISG56/IFIT1 Gene Family. Journal of Interferon & Cytokine Research, 31(1), 71–78. https://doi.org/10.1089/jir.2010.0101
2. Abbas, Y. M., Pichlmair, A., Górna, M. W., Superti-Furga, G., & Nagar, B. (2013). Structural basis for viral 5′-PPP-RNA recognition by human IFIT proteins. Nature, 494(7435), 60–64. https://doi.org/10.1038/nature11783
3. Rehwinkel, J., & Reis E Sousa, C. (2013). Targeting the viral Achilles’ heel: Recognition of 5′-triphosphate RNA in innate anti-viral defence. Current Opinion in Microbiology, 16(4), 485–492. https://doi.org/10.1016/j.mib.2013.04.009
4. Zheng, C., Zheng, Z., Zhang, Z., Meng, J., Liu, Y., Ke, X., Hu, Q., & Wang, H. (2015). IFIT5 positively regulates NF-κB signaling through synergizing the recruitment of IκB kinase (IKK) to TGF-β-activated kinase 1 (TAK1). Cellular Signalling, 27(12), 2343–2354. https://doi.org/10.1016/j.cellsig.2015.08.018
5. Lo, U.-G., Pong, R.-C., Yang, D., Gandee, L., Hernandez, E., Dang, A., Lin, C.-J., Santoyo, J., Ma, S., Sonavane, R., Huang, J., Tseng, S.-F., Moro, L., Arbini, A. A., Kapur, P., Raj, G. V., He, D., Lai, C.-H., Lin, H., & Hsieh, J.-T. (2019). IFNγ-Induced IFIT5 Promotes Epithelial-to-Mesenchymal Transition in Prostate Cancer via miRNA Processing. Cancer Research, 79(6), 1098–1112. https://doi.org/10.1158/0008-5472.CAN-18-2207
6. Huang, J., Lo, U.-G., Wu, S., Wang, B., Pong, R.-C., Lai, C.-H., Lin, H., He, D., Hsieh, J.-T., & Wu, K. (2019). The roles and mechanism of IFIT5 in bladder cancer epithelial–mesenchymal transition and progression. Cell Death & Disease, 10(6), 437. https://doi.org/10.1038/s41419-019-1669-z
Back to Top