The cystic fibrosis transmembrane conductance regulator (CFTR) is a chloride ion channel belonging to the ATP-binding cassette (ABC) transporter superfamily. Unlike typical ABC transporters, which use ATP hydrolysis to drive substrate movement across membranes, CFTR uses ATP primarily to regulate channel gating, the opening and closing of the pore that allows chloride flux. Dysfunction of CFTR causes cystic fibrosis (CF), the most common lethal autosomal recessive disorder in caucasian populations. The most prevalent mutation, ΔF508, occurs in the first nucleotide-binding domain (NBD1) and disrupts folding, stability, and trafficking of the protein .
NBD1 is one of two cytosolic domains that bind ATP. Because NBD1 contains many disease-associated mutations and participates directly in ATP-dependent gating, elucidating its structure is essential for understanding how CFTR functions and how mutations disrupt the channel activity. Lewis et al. (2004) determined crystal structures of mouse NBD1 in several nucleotide-bound and unliganded states, providing insight into CFTR’s unique regulatory features and the molecular basis of CF-causing mutations.
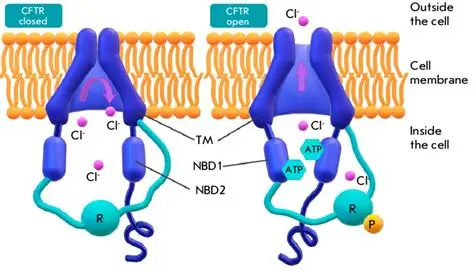
Figure 1. Schematic of the cystic fibrosis transmembrane conductance regulator (CFTR) in the closed (left) and open (right) states. Phosphorylation of the regulatory (R) domain and ATP binding to nucleotide-binding domains NBD1 and NBD2 promote channel opening, allowing chloride (Cl⁻) ions to pass through the transmembrane pore across the cell membrane.
II. General Structure
CFTR NBD1 adopts the canonical fold shared by ABC transporter nucleotide-binding domains but includes several unique structural features that distinguish it from typical examples. Like other ABC NBDs, it consists of two major subdomains: (1) an F1-type ATP-binding core containing the Walker A
and Walker B
motifs and (2) the ABC-specific α-helical subdomain , which includes the LSGGQ signature sequence in NBD2 but not in NBD1. Despite this similarity, NBD1 includes a prominent regulatory insertion between β-strands S1 and S2 , composed of short helices (H1b and H1c)
and a flexible loop (ommited) which suggest that the flexible loop is not vital for NMD1 function.
A second distinctive feature is the regulatory extension at the C-terminus, where an extended α-helix (H9b) contains two PKA phosphorylation sites (Ser660 and Ser670). These sites, along with Ser422 in the insertion region, become ordered upon phosphorylation.
The overall core structure of NBD1 remains strikingly similar across nucleotide-free, ADP-bound, and ATP-bound crystal forms, implying that binding does not induce major conformational changes within isolated NBD1.
III. ATP Binding
ATP binds within a conserved pocket formed by the Walker A and Walker B motifs of NBD1, and is coordinated through hydrogen bonding and Mg2+ induced bonds with Lys464 and Thr465 (Walker A), Asp572 (Walker B), and Gln493 (the Q-loop) .
However, NBD1 is catalytically inactive on its own. The catalytic glutamate normally found in ABC transporters is replaced by Ser573, and the usual H-loop histidine is replaced by serine at residue 605, eliminating residues required for activating water for nucleophilic attack on the γ-phosphate .
The nucleotide also assumes an unusual conformation. The adenine base does not stack against a conserved aromatic residue; instead, it is contacted laterally by Phe430 and lies adjacent to Trp401, Leu409 makes contact by approaching from the opposite side of the ring .
This altered geometry reflects displacement produced by the regulatory insertion.
IV. Domain Interactions
NBD1 normally interacts with NBD2 to form an asymmetric ATP-sandwich dimer, where ATP bound at the NBD1–NBD2 interface participates in gating. However, structural modeling using the crystallized NBD1 structure revealed severe steric clashes between NBD1’s regulatory insertion and the predicted NBD2 interface.
Elevated B-factors in the insertion (residues 405–436) and in helices H8–H9b indicate inherent flexibility .
Phosphorylation may facilitate these rearrangements, and all mapped phosphorylation sites cluster in these flexible regions. NBD1 also interacts with MSD1 (membrane-spanning domain 1). Structural modeling predicts that the NBD1 interface contacting MSD1 includes several hydrophobic residues, including the highly significant Phe508
.
V. Disease-Relevant Mutations
Many cystic-fibrosis-causing mutations map to NBD1, and the Lewis et al. structure explains their pathogenicity. The most common mutation, ΔF508, deletes phenylalanine 508 at the junction of helices H3 and H4 .
Although Phe508 forms only modest intradomain contacts, the structure reveals that it lies directly within the predicted interface between NBD1 and MSD1. Thus, loss of Phe508 disrupts interdomain packing rather than ATP binding.
Other CF mutations cluster on the α-subdomain or near the NBD1–NBD2 interface, where they may disturb interdomain rearrangements essential for gating. Still others (e.g., G551D, S549R) affect residues near the canonical ATP-binding motifs, compromising nucleotide coordination despite NBD1’s limited hydrolytic capacity. Altogether, the Lewis et al. structure shows that CF-causing mutations in NBD1 often impair structural stability, interdomain contacts, or nucleotide-dependent conformational transitions.
VI. References
Annereau JP, Ko YH, Pedersen PL (2003) Cystic fibrosis transmembrane conductance regulator: the NBF1+R (nucleotide‐binding fold 1 and regulatory domain) segment acting alone catalyses a Co2+/Mn2+/Mg2+‐ATPase activity markedly inhibited by both Cd2+ and the transition‐state analogue orthovanadate. Biochem J 371: 451–462
Bobadilla JL, Macek Jr M, Fine JP, Farrell PM (2002) Cystic fibrosis: a worldwide analysis of CFTR mutations—correlation with incidence data and application to screening. Hum Mutat 19: 575–606
Lewis, H.A. et al. 2004. Structure of nucleotide‐binding domain 1 of the cystic fibrosis transmembrane conductance regulator. The EMBO Journal 23:282–293.
Back to Top