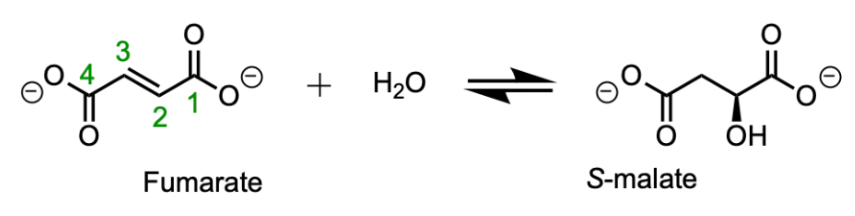
Figure 2. The fumarate to malate reaction and its reverse reaction are catalyzed by fumarase.
II. General Structure
The class II fumarases, to which this yeast enzyme belongs, share common structural motifs. All are
proteins containing three domains labeled
. The central domain is
which is composed of a five helix bundle
. In the tetrameric structure, this core becomes a 20 alpha helix
. These helices vary in length from 25-35 residues and most have some curvature to accommodate tight packing in the hydrophobic core. While the tetrameric protein is mostly alpha-helical, some regions do form beta sheets. The N-terminal
region is D1 and D3 is located at the C-terminus
of the polypeptide. Regions D1 and D3 form the caps on either side of the helical domain. These domains come together to form the active site in the tetrameric enzyme. Specific amino acid residues are also common to these proteins. The three consensus sequences:
H154-T171,
I206-E225 and
P337-E356 define this family of enzymes.
III. Active Site
Fumarase contains a multi-subunit active site where the three conserved sequences mentioned above
. This space is a crevice between domains D1 and D2. Keep in mind that with four subunits, this enzyme fits to form four active sites, adding to its efficiency. This crystal structure was generated with meso-tartrate in the active site, a competitive inhibitor of this enzyme which mimics the native substrate due to its size and charged carboxylate groups (ligand not shown). Fumarate, the enzyme’s typical substrate, is a four-carbon dicarboxylic acid featuring a trans double bond at C2. In the reversible reaction catalyzed by the enzyme a hydroxyl group is introduced in a stereoselective manner through hydration of the double bond. The precise catalytic mechanism is still an open question. The forward reaction (fumarate → S-malate) is better understood than the reverse reaction. The reverse, dehydration, reaction is impressive due to the high pKa of the beta proton eliminated without the use of cofactors like NAD or NADPH.
In this paper the authors speculate that a carbanion intermediate exists during the conversion of S-malate to fumarate, arising from migration of electrons participating in the double bond upon addition of -OH. The
K349c and N351c may stabilize this carbanion. A water molecule was bound in the active sites of both native and recombinant yeast fumarase. This molecule is likely key for catalysis since its activation by a general base forms the requisite hydroxyl anion which attacks C2 in the forward reaction. The water molecule forms hydrogen bonds with
: H213, S124, N166 and T126.
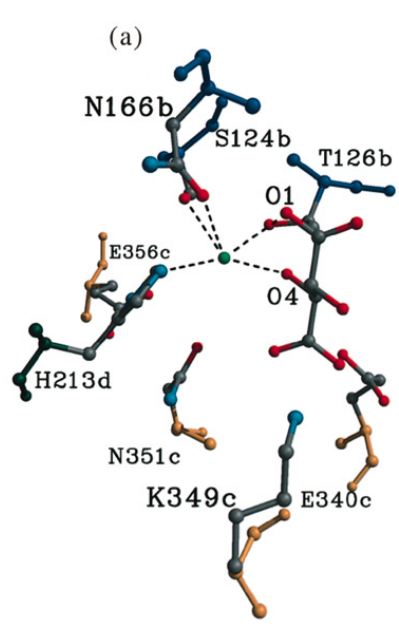
Figure 3. Molecular image of four residues (H213, N166, T126, and S124) that interact with a highly conserved water molecule in the active site. This water molecule, though not included in the crystalized structure, is the attacking water molecule in this reaction. Residues H213 and E356 are involved in its activation.
A charge relay chain spanning
and H213 is predicted to activate this water molecule during the reaction. This histidine has a dual function as both a catalytic base to activate water, allowing removal of the C3 proton of S-malate and, when charged, it provides a binding site for the C1 carboxylate of the substrate. Crucially, this water is positioned closest to the C3 proton of meso-tartarate which is eliminated in the formation of fumarate–suggesting it acts as the catalytic base in this reaction (6).
Several facts about the process are well supported by experimental evidence, however. We know the enzyme/H2O/fumarate complex decays to malate after exchange of water faster than fumarate can diffuse out of the active site due to the high rate of labeling in malate recovered after brief conversion to fumarate in 18O water (3). The rapid incorporation of the beta proton abstracted from malate into fumarate and other analogs indicates slow recycling of the enzyme conjugate acid group formed during the reaction. This means the free proton pulled off of malate in the dehydration reaction ended up on the carboxylate group of the substrate before it could return to the general base from the enzyme. When fumarase was saturated with deuterium (D2O) the maximum rate of the forward reaction was reduced by 60%, for the reverse reaction this number was 33% (4). This suggests that the heavier hydrogen isotope is more deleterious to the enzyme recycling process than the reaction chemistry itself.
Fumarase generally exhibits negative cooperativity, meaning binding of a substrate to one subunit of the tetrameric protein reduces the affinity of other subunits for the same ligand. At sub 1mM fumarate concentrations the enzyme exhibits simple Michaelis-Menten kinetics, meaning the reaction rate steadily increases with increasing substrate concentration. At higher concentrations of fumarate (0.001– 0.033 M), allosteric activation begins and the affinity of each subunit for the substrate increases. However, as the concentration increases to 0.1 M or higher apparent inhibition occurs (5).
IV. A Second Dicarboxylic Acid Binding Site
In eukaryotes, the tetrameric fumarase complex includes additional sites capable of binding both reactants and inducing allosteric changes to enzyme activity. A dicarboxylic acid binding site was discovered in previous crystallographic studies of the prokaryotic isozyme. This site was 12 Å from the active site, toward the N-terminus at the end of a short π-helix. This information allowed Weaver et. al. to investigate the comparable location in NY-fumarase for an analogous binding site.
, a short π-helical segment between residues Q152 and C159 is present in NY-fumarase. Since residues L144 to Q152 are not present in this crystal structure, the observable π-helix comprises V153 to C159.
residues K151, H154, N156 and N157 in this helix might be involved in salt-briding interactions with the dicarboxylic acid substrates. It is not clear how binding interaction would impact enzyme turnover; the authors suggest mutation studies to connect binding in this site with allosteric regulation.
V. References
Meléndez-Hevia, E.; Waddell, T.G.; Cascante, M. The puzzle of the Krebs citric acid cycle: Assembling the pieces of chemically feasible reactions, and opportunism in the design of metabolic pathways during evolution. Journal of Molecular Evolution 1996 43, 293–303. DOI: 10.1007/BF02338838. Mescam, M.; Vinakota, K. C.; Beard, D. A. Identification of the Catalytic Mechanism and Estimation of Kinetic Parameters for Fumarase. Journal of Biological Chemistry 2011 286 (24), 21100-21109. DOI: 10.1074/jbc.M110.214452.
Rose, I. A. How fumarase recycles after the malate --> fumarate reaction. Insights into the reaction mechanism. Biochemistry. 1998 37 (51), 17651-8. DOI: 10.1021/bi9821521.
Thomson, J. F. Fumarase activity in D2O. Archives of Biochemistry and Biophysics. 1960 90 (1), 1-6. DOI: 10.1016/0003-9861(60)90602-0.
Weaver, T.; Lees, M.; Zaitsev, V.; Zaitseva, I.; Duke, E.; Lindley, P.; McSweeny, S.; Svensson, A.; Keruchenko, J.; Keruchenko, I.; Gladilin, K.; Banaszak, L. Crystal structures of native and recombinant yeast fumarase. Journal of Molecular Biology 1998 280 (3), 431-442. DOI: 10.1006/jmbi.1998.1862.
Back to Top