Mus musculus Microphthalmia-Associated Transcription Factor (MITF)
Damaris Darkwah '27 and Shoshana Marcus '28
View Type:
Contents:
I. Introduction
The microphthalmia-associated transcription factor (MITF) is a member of the basic helix-loop-helix zipper (bHLHZip) transcription factor protein family. MITF is a master transcription factor for the developement of melanocytes, the cells responsible for synthesizing melanin, which determines skin and eye color. It plays a crucial role in regulating their development, survival, and differentiation. MITF is also essential for the proper development of several cell lineages, including osteoclasts, retinal pigment epithelial (RPE) cells, and mast cells. Additionally, MITF is involved in various pathophysiological processes, including allergic responses and autophagy. It can bind to protein as a homodimer or as a heterodimer with other members of the TFE protein family. The MITF protein consists of amino acids.
Studying MITF is important because it functions as a lineage-specific survival oncogene in malignant melanoma. In humans, mutations in MITF can lead to two pigmentation and deafness syndromes known as Waardenburg syndrome type 2A and Tietz syndrome. Similar effects, including poor eye development, are also observed in mice with mutations in the Mitf gene.
II. General Structure
The MITF protein consists of a basic/binding region responsible for specific DNA binding, a helix-loop-helix region, and a leucine zipper domain . The helix-loop-helix and leucine zipper are critical for protein-protein interactions, allowing MITF to form a functional dimer. The leucine zipper domain has an unusual kink that limits dimerization with other basic helix-loop-helix zipper (bHLHZ) transcription factors. As a bHLHZip protein, MITF binds to a palindromic CACGTG E-box . However, unlike other bHLHZip transcription factors, MITF also interacts with the M-box sequence TCATGTG, which has a flanking thymidine in the -4 position at the 5' end. These unusual structural features of MITF explain its distinct role as a master transcription factor.
III. Leucine Zipper
The leucine zipper consists of two protomers that extend from the second helix-loop-helix structure, denoted as ⍺2 and ⍺2';. This region contains five heptad repeats, labeled 0 through IV.
The first protomer, ⍺2, is made up of ⍺2a and ⍺2b . The ⍺2a helix terminates after the first recognizable heptad repeat, referred to as repeat 0 (see Figure 1). Within this heptad, a cluster of intermolecular is formed by the side chains of Tyr253 and Gln258
and the main chain carbonyl group of residue 230 from the C terminus of the first DNA-binding helix.
Following this region is a marked
which determines determization specifity. It is caused by a three residue insertion in the heptad repeat register at Arg259-Gln261. This is unique to the bHLHZip transcription factors of the TFE family. This insertion is neccessary for heterodimerization binding.
After the kink, the ⍺2b helix extends over the remaining heptad repeats I through IV (see Figure 1).
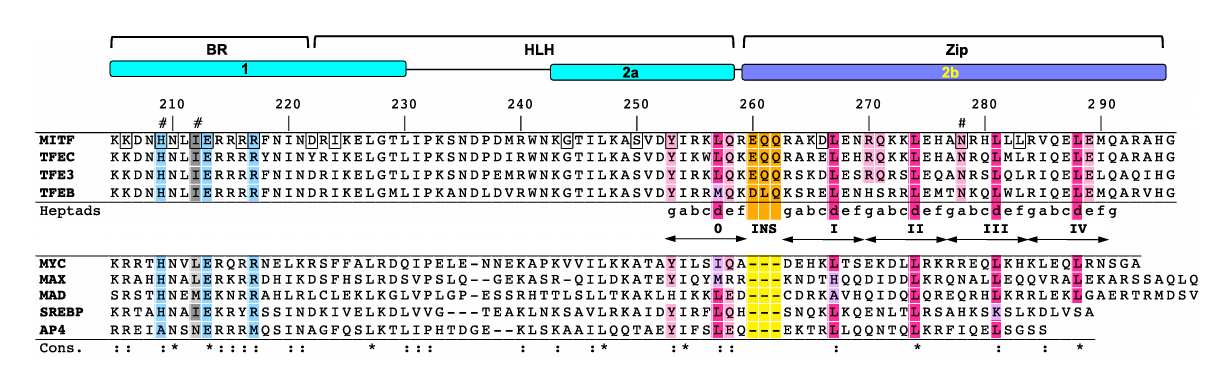
Figure 1. Multiple sequence alignment of MITF, members of the TFE family, and canonical bHLHZip transcription factors. Labels 0-IV represent the heptad repeat, with the three residue insertion highlighted in orange. (Pogenberg et al.)
The , ⍺2'
, consists of a long helix with almost 15 turns. The kink creates a gap in the structure due to the absence of the typical leucine zipper interaction.
IV. Helix-Loop-Helix
The HLH
consists of two ⍺ helices, 1 and 2; the N-terminal part of the
extended ⍺ helix constitutes the basic region and provides most of the DNA major groove contacts. The following HLH domain is composed of the C-terminal part of the first helix and the N-terminal part of the , separated by a .
V. DNA Binding
Binding is mediated by His209 , Glu213, Ile212, and Arg217, which provide key base-specific interactions. Arg217 can bind to the Guanine in the +1 position only in the Mbox. Glu213 can bind to the Cytosince in the -3 position in both the Mbox and Ebox motif. The ability of MITF to specifically recognize E-box and M-box sequences is primarily driven by the residues His209 and Ile212. The His209 imidazole ring of both MITF protomers points symmetrically into the bottom of the M-box major groove, nearly equidistant to both DNA strands. This allows for with bases in both strand in positions -4/+3. In the Mbox, Ile212 also interacts with the thymidine base from the -4 position. This interaction is and is crucial for target gene selection of M-box-type promoters.
.
VI. Mutations
Because mammalian coat-color mutants are easy to identify and study, numerous spontaneous mutations in Mitf have been discovered across many species. When Mitf is mutated in mice, the phenotype is characterized by bone hyperdensity, small eyes, deafness, and absence of pigment in eyes and skin. Non-functional mutation of Mitf leads to a coat that lacks color due to lack of melanocytes. Heterozygotes usually have white or pale spotting. These spots have been associated with mutations within the DNA binding regions. An example of such a mutation is As previously mentioned, Ile212 is very important Mbox binding, therefore this mutation results substantial reduction of Mbox binding. COMMAD (coloboma, osteopetrosis, microphthalmia, macrocephaly, albinism, and deafness) syndrome is a human genetic disorder associated MITF biallelic mutants for example compound heterozygosity for which also impairs DNA binding.
VII. References
Pogenberg V, Ogmundsdóttir MH, Bergsteinsdóttir K, Schepsky A, Phung B, Deineko V, Milewski M, Steingrímsson E, Wilmanns M. Restricted leucine zipper dimerization and specificity of DNA recognition of the melanocyte master regulator MITF. Genes Dev. 2012 Dec 1;26(23):2647-58. doi: 10.1101/gad.198192.112. PMID: 23207919; PMCID: PMC3521630.
Levy, C., Khaled, M., & Fisher, D. E. (2006). MITF: master regulator of melanocyte development and melanoma oncogene. Trends in molecular medicine, 12(9), 406–414. https://doi.org/10.1016/j.molmed.2006.07.008
Lee, A., Lim, J., & Lim, J. S. (2024). Emerging roles of MITF as a crucial regulator of immunity. Experimental & molecular medicine, 56(2), 311–318. https://doi.org/10.1038/s12276-024-01175-5
Liu, Z., Chen, K., Dai, J. et al. A unique hyperdynamic dimer interface permits small molecule perturbation of the melanoma oncoprotein MITF for melanoma therapy. Cell Res 33, 55–70 (2023). https://doi.org/10.1038/s41422-022-00744-5
Jón H Hallsson, Benedikta S Haflidadóttir, Chad Stivers, Ward Odenwald, Heinz Arnheiter, Francesca Pignoni, Eiríkur Steingrímsson, The Basic Helix-Loop-Helix Leucine Zipper Transcription Factor Mitf Is Conserved in Drosophila and Functions in Eye Development, Genetics, Volume 167, Issue 1, 1 May 2004, Pages 233–241, https://doi.org/10.1534/genetics.167.1.233
Aksan I, Goding CR. Targeting the microphthalmia basic helix-loop-helix-leucine zipper transcription factor to a subset of E-box elements in vitro and in vivo. Mol Cell Biol. 1998 Dec;18(12):6930-8. doi: 10.1128/MCB.18.12.6930. PMID: 9819381; PMCID: PMC109276.
Udono, T., Yasumoto, K., Takeda, K., Amae, S., Watanabe, K., Saito, H., Fuse, N., Tachibana, M., Takahashi, K., Tamai, M., & Shibahara, S. (2000). Structural organization of the human microphthalmia-associated transcription factor gene containing four alternative promoters. Biochimica et biophysica acta, 1491(1-3), 205–219. https://doi.org/10.1016/s0167-4781(00)00051-8
Back to the top