Gambier COVID Virus, Influenza
Virus and
Environmental Air Quality
This site reports recent
environmental data of interest to Gambier, Ohio and Kenyon
College. The Village of Gambier monitors wastewater for levels
of RNA from dead coronavirus
SARS-CoV-2, the cause of COVID disease; and for influenza virus. Samples are from the Village of
Gambier Wastewater Treatment Plant. From August, 2022, viral
RNA N2 has been measured and reported by the Ohio Department of
Health.
For questions or weekly updates contact: slonczewski [at] kenyon.edu
For archival COVID data see: 2020 - 2023 Archival COVID dataSpring 2025
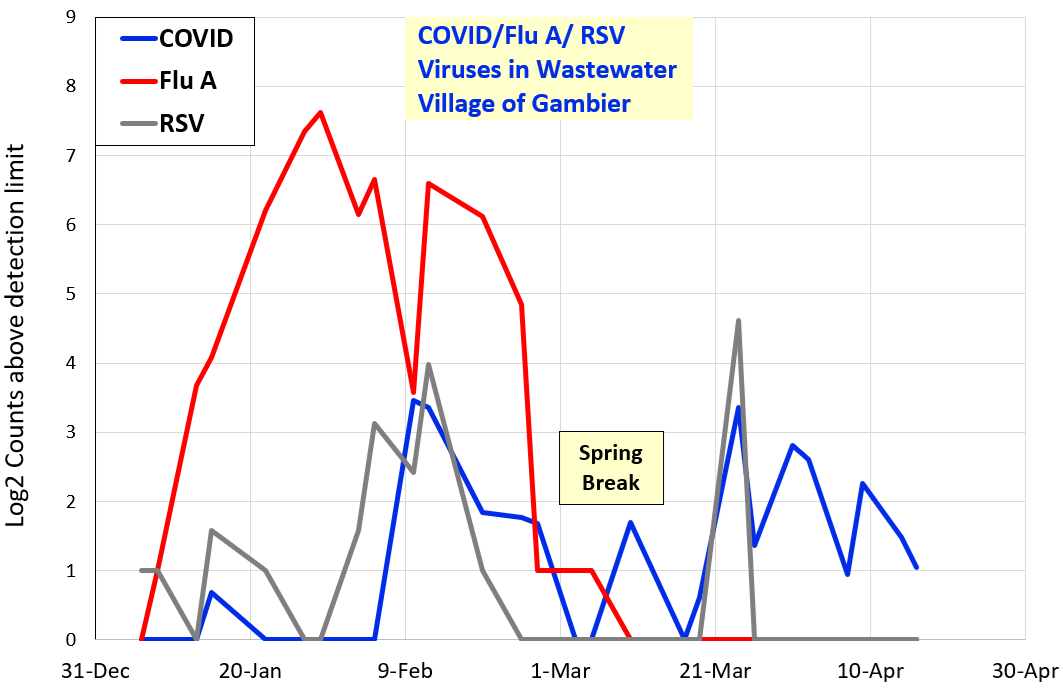
Fall 2024
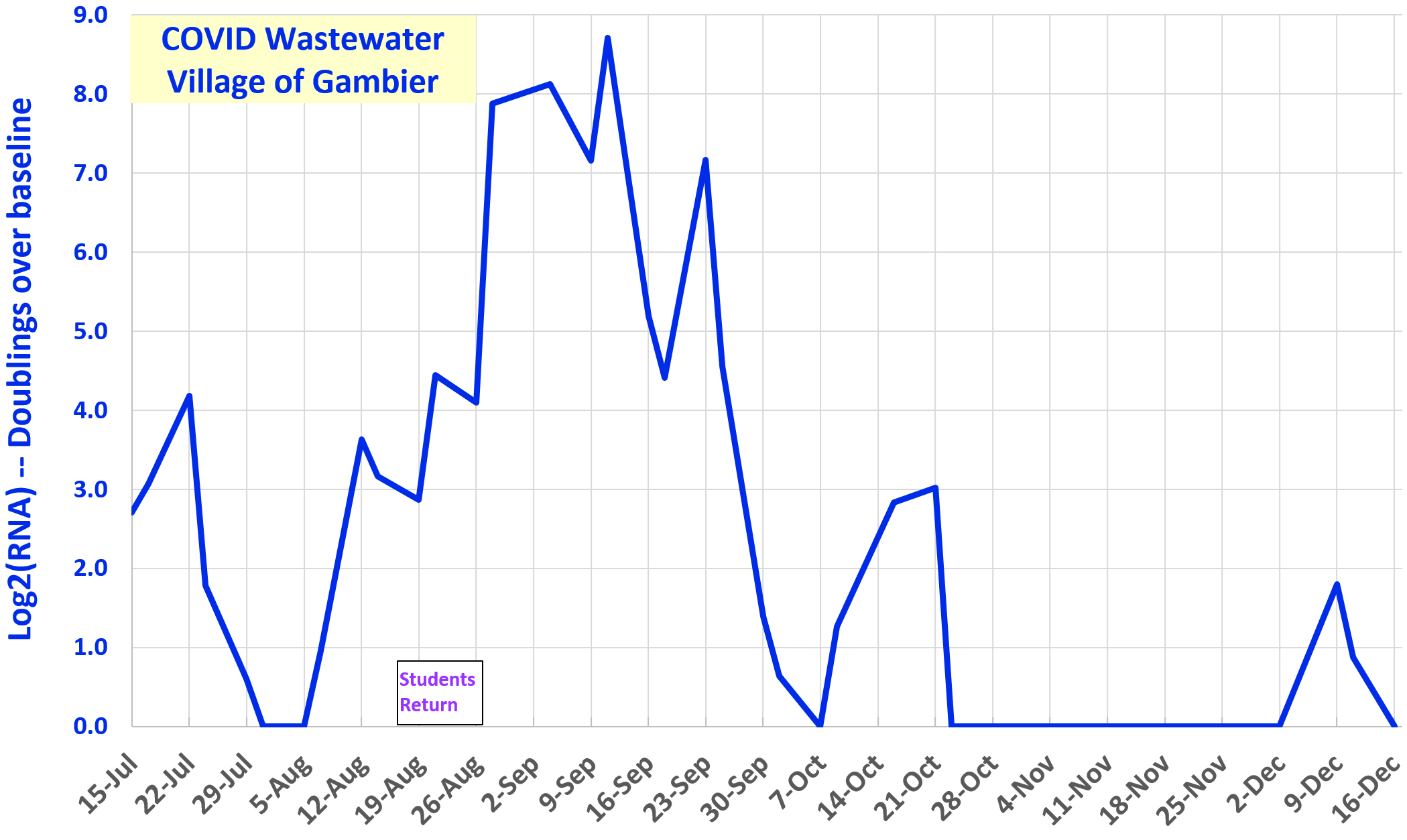
Spring 2024
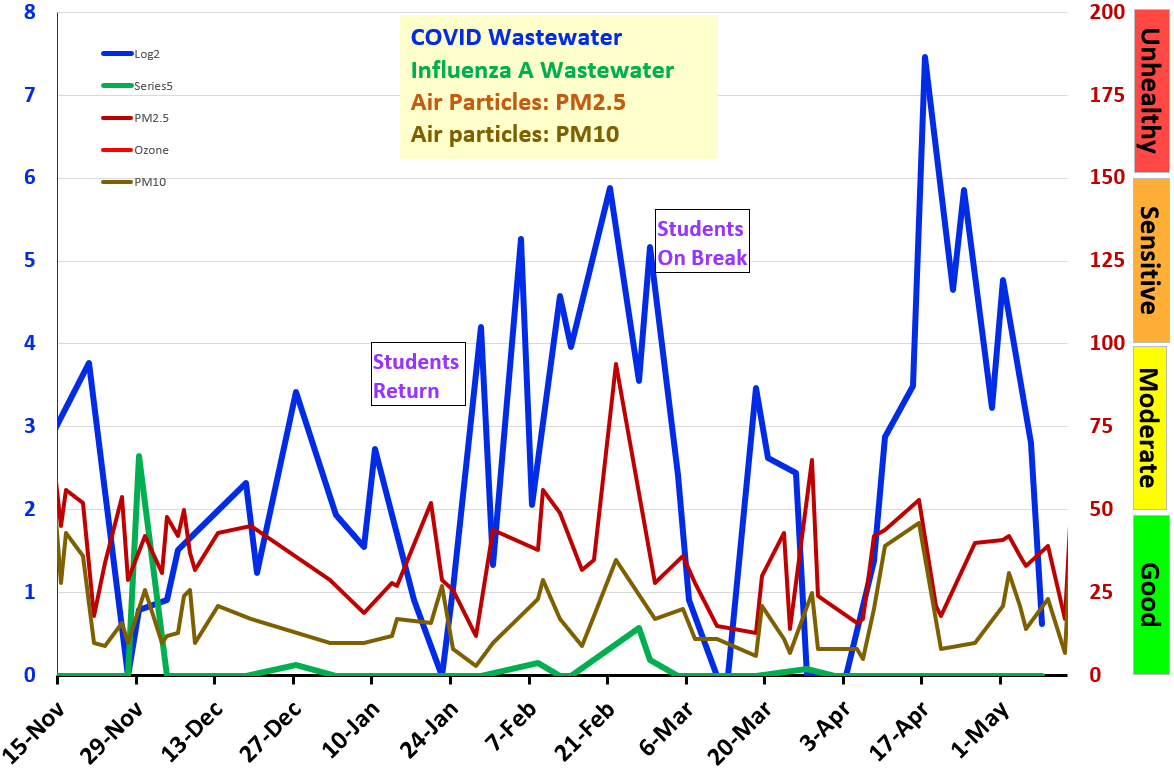
Explanation:
COVID signal is the log2 concentration of an RNA gene from SARS-CoV-2 virus, in a sample from the Village of Gambier wastewater treatment plant. The log2 signal is now normalized to the limit of detection (LOD).
Influenza virus RNA is now tested in Gambier wastewater from the same samples as COVID. Influenza RNA testing is highly experimental, but rise and fall of signal is predicted to show rise and fall of case numbers.
PM2.5 and PM10 are sizes of microscopic particles from smoke (and other sources) reported to AirNow.gov.
Sources:
https://coronavirus.ohio.gov/dashboards/other-resources/wastewater
https://www.airnow.gov/